Methods and Parameters for Feature Detection: Part 2
In this notebook, we will contninue to look in detail at tobac’s feature detection and examine the remaining parameters.
We will treat:
[1]:
import matplotlib.pyplot as plt
import numpy as np
import xarray as xr
%matplotlib inline
import seaborn as sns
sns.set_context("talk")
import warnings
warnings.filterwarnings("ignore")
[2]:
import tobac
import tobac.testing
Object Erosion Parameter n_erosion_threshold
To understand this parameter we have to look at one varibale of the feature-Datasets we did not mention so far: num
The value of num for a specific feature tells us the number of datapoints exceeding the threshold. n_erosion_threshold reduces this number by eroding the mask of the feature on its boundary. Supose we are looking at the gaussian data again and we set a treshold of 0.5. The resulting mask of our feature will look like this:
[3]:
x = np.linspace(-2, 2)
y = np.linspace(-2, 2)
xx, yy = np.meshgrid(x, y)
exp = np.exp(-(xx**2 + yy**2))
gaussian_data = np.expand_dims(exp, axis=0)
threshold = 0.5
mask = gaussian_data > threshold
mask = mask[0]
plt.figure(figsize=(8, 8))
plt.imshow(mask)
plt.show()
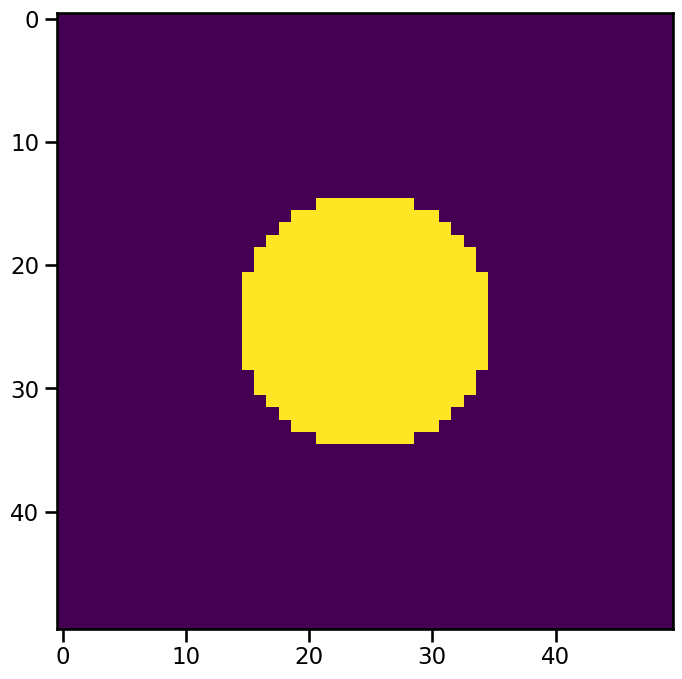
The erosion algorithm used by tobac is imported from skimage.morphology:
[4]:
from skimage.morphology import binary_erosion
Applying this algorithm requires a quadratic matrix. The size of this matrix is provided by the n_erosion_threshold parameter. For a quick demonstration we can create the matrix by hand and apply the erosion for different values:
[5]:
fig, axes = plt.subplots(ncols=3, figsize=(14, 10))
im0 = axes[0].imshow(mask)
axes[0].set_title(r"$\mathtt{n\_erosion\_threshold} = 0$", fontsize=14)
n_erosion_threshold = 5
selem = np.ones((n_erosion_threshold, n_erosion_threshold))
mask_er = binary_erosion(mask, selem).astype(np.int64)
im1 = axes[1].imshow(mask_er)
axes[1].set_title(r"$\mathtt{n\_erosion\_threshold} = 5$", fontsize=14)
n_erosion_threshold = 10
selem = np.ones((n_erosion_threshold, n_erosion_threshold))
mask_er_more = binary_erosion(mask, selem).astype(np.int64)
im2 = axes[2].imshow(mask_er_more)
axes[2].set_title(r"$\mathtt{n\_erosion\_threshold} = 10$", fontsize=14)
[5]:
Text(0.5, 1.0, '$\\mathtt{n\\_erosion\\_threshold} = 10$')
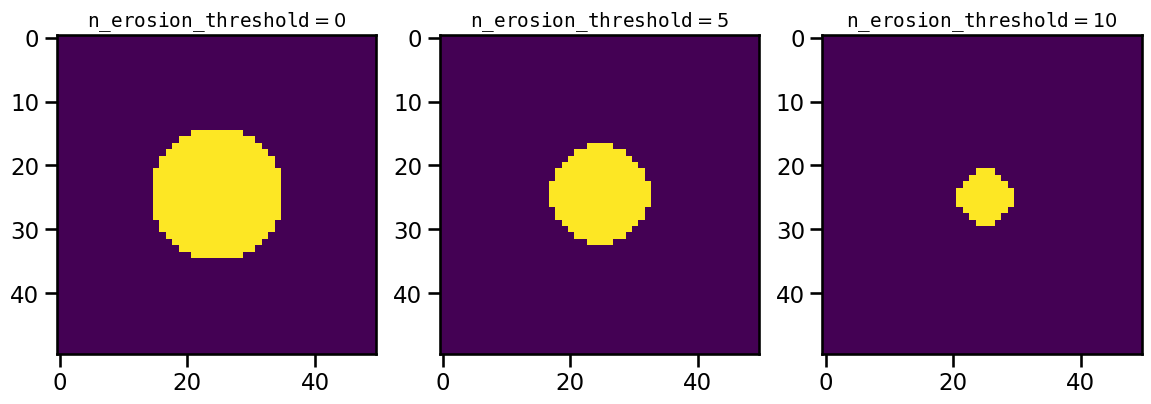
This means by using increasing values of n_erosion_threshold for a feature detection we will get lower values of num, which will match the number of True-values in the masks above:
[6]:
%%capture
date = np.datetime64("2022-04-01T00:00")
input_data = xr.DataArray(
data=gaussian_data, coords={"time": np.expand_dims(date, axis=0), "y": y, "x": x}
)
dxy = input_data["x"][1] - input_data["x"][0]
features = tobac.feature_detection_multithreshold(
input_data, dxy, threshold, n_erosion_threshold=0
)
features_eroded = tobac.feature_detection_multithreshold(
input_data, dxy, threshold, n_erosion_threshold=5
)
features_eroded_more = tobac.feature_detection_multithreshold(
input_data, dxy, threshold, n_erosion_threshold=10
)
[7]:
features["num"][0]
[7]:
332
[8]:
mask.sum()
[8]:
332
[9]:
features_eroded["num"][0]
[9]:
188
[10]:
mask_er.sum()
[10]:
188
[11]:
features_eroded_more["num"][0]
[11]:
57
[12]:
mask_er_more.sum()
[12]:
57
This can be used to simplify the geometry of complex features.
Minimum Object Size Parameter n_min_threshold
With n_min_threshold parameter we can exclude smaller features by setting a minimum of datapoints that have to exceed the threshold for one feature. If we again detect the three blobs and check their num value at frame 50, we can see that one of them contains fewer pixels.
[13]:
data = tobac.testing.make_sample_data_2D_3blobs(data_type="xarray")
plt.figure(figsize=(8, 8))
plt.imshow(data[50])
plt.show()
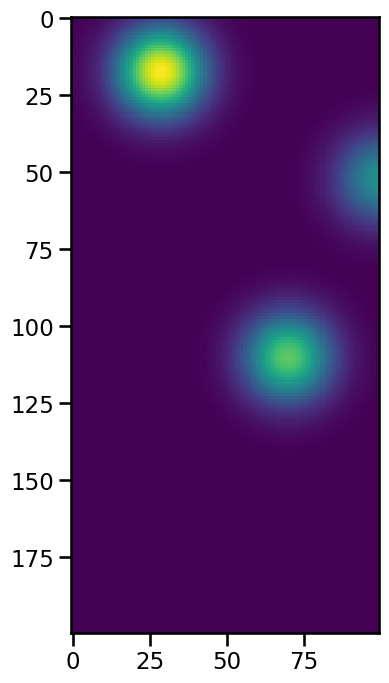
[14]:
%%capture
threshold = 9
dxy, dt = tobac.utils.get_spacings(data)
features = tobac.feature_detection_multithreshold(data, dxy, threshold)
[15]:
features.where(features["frame"] == 50)["num"].dropna()
[15]:
60 501.0
61 30.0
62 325.0
Name: num, dtype: float64
Obviously, the feature with only 30 datapoints is the rightmost feature that has almost left the imaging area. If we now use an n_min-threshold above or equal to 30, we will not detect this small feature:
[16]:
%%capture
features = tobac.feature_detection_multithreshold(
data, dxy, threshold, n_min_threshold=30
)
[17]:
features.where(features["frame"] == 50)["num"].dropna()
[17]:
60 501.0
61 325.0
Name: num, dtype: float64
Noisy data can be cleaned using this method by ignoring smaller features of no significance or, as here, features that leave the detection range.
Minimum Object Pair Distance min_distance
Another way of getting rid of this specific feature is the min_distance parameter. It sets a minimal distance of our features. Lets detect the features as before:
[18]:
%%capture
data = tobac.testing.make_sample_data_2D_3blobs(data_type="xarray")
threshold = 9
dxy, dt = tobac.utils.get_spacings(data)
features = tobac.feature_detection_multithreshold(data, dxy, threshold)
A quick look at frame 50 tells us the found features and their indices:
[19]:
n = 50
mask = features["frame"] == n
features_frame = features.where(mask).dropna()
features_frame.index.values
[19]:
array([60, 61, 62])
Notice that to_dataframe() was used to convert the Dataset to a pandas dataframe, which is required to use the calculate_distance function of the analysis module. The distances bewteen our features are:
[20]:
tobac.analysis.calculate_distance(
features_frame.loc[60], features_frame.loc[61], method_distance="xy"
)
[20]:
77307.67873610597
[21]:
tobac.analysis.calculate_distance(
features_frame.loc[62], features_frame.loc[61], method_distance="xy"
)
[21]:
64289.48419281164
[22]:
tobac.analysis.calculate_distance(
features_frame.loc[62], features_frame.loc[60], method_distance="xy"
)
[22]:
101607.57596570993
With this knowledge we can set reasonable values for min_distance to exclude the small feature:
[23]:
min_distance = 70000
and perform the feature detection as usual:
[24]:
%%capture
data = tobac.testing.make_sample_data_2D_3blobs(data_type="xarray")
thresholds = [10]
dxy, dt = tobac.utils.get_spacings(data)
features = tobac.feature_detection_multithreshold(
data, dxy, thresholds, min_distance=min_distance
)
n = 50
mask = features["frame"] == n
Plotting the result shows us that we now exclude the expected feature.
[25]:
fig, ax1 = plt.subplots(ncols=1, figsize=(8, 8))
ax1.imshow(data.data[50], cmap="Greys")
im1 = ax1.scatter(
features.where(mask)["hdim_2"], features.where(mask)["hdim_1"], color="red"
)
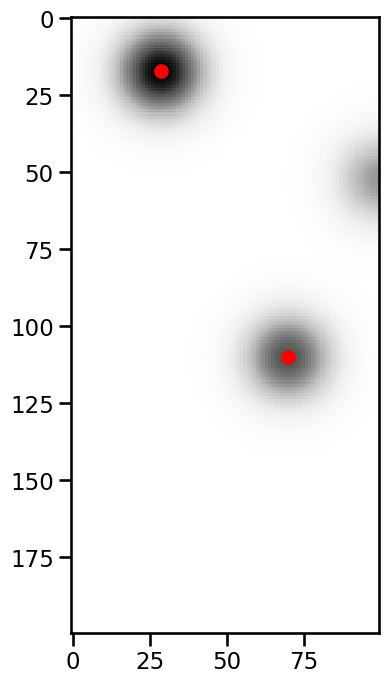
If the features have the same threshold, tobac keeps the feature with the larger area. Otherwise the feature with the higher treshold is kept.